Preprints
(# indicates equal contributions, * indicates corresponding authors)
2024
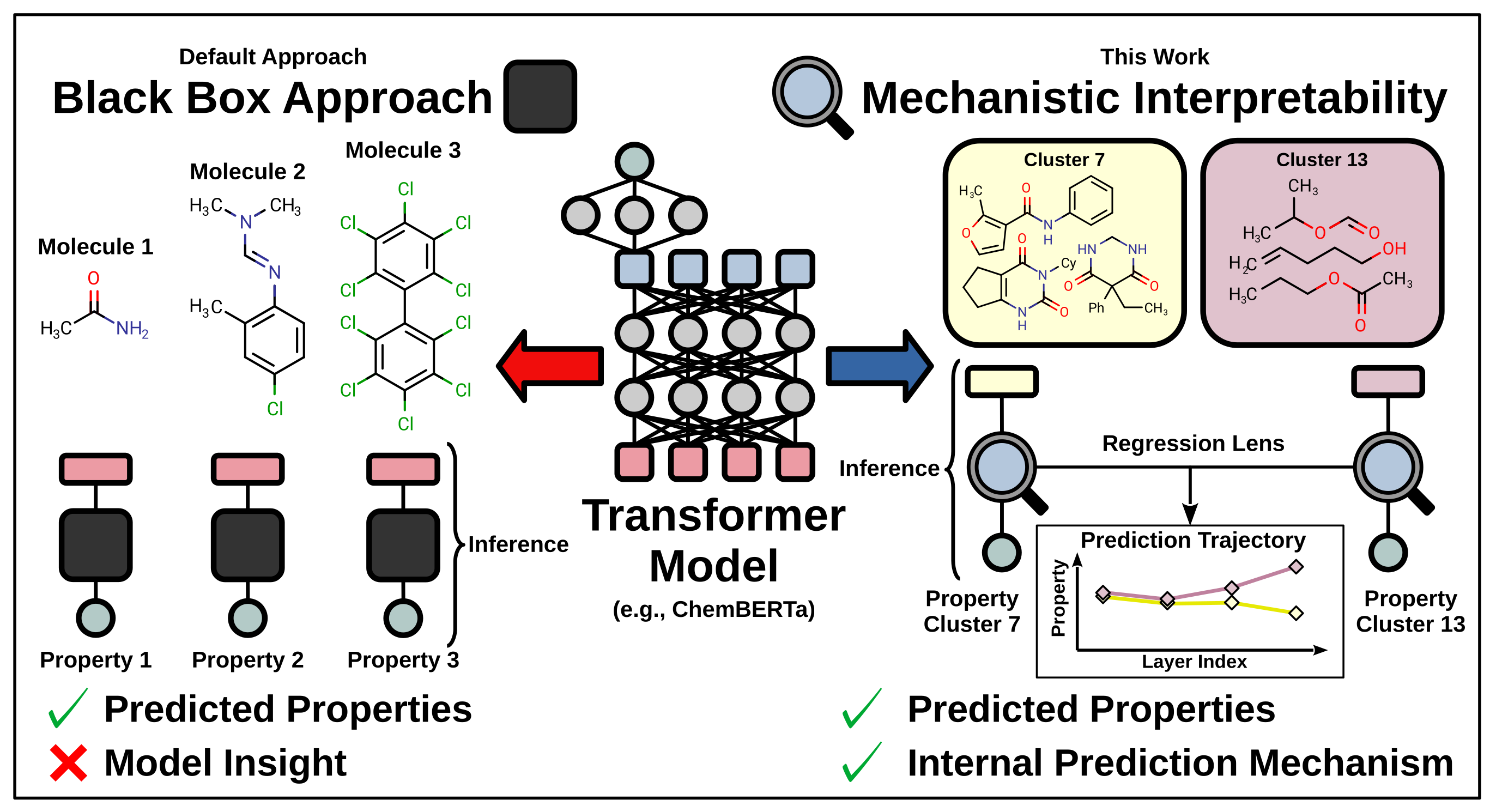
24. "Uncovering Internal Prediction Mechanisms of Transformer-Based Chemical Foundation Models"
Artificial Design
A. Müller, J. Cardenas-Cartagena, R. Pollice*
ChemRxiv 2025.
2024
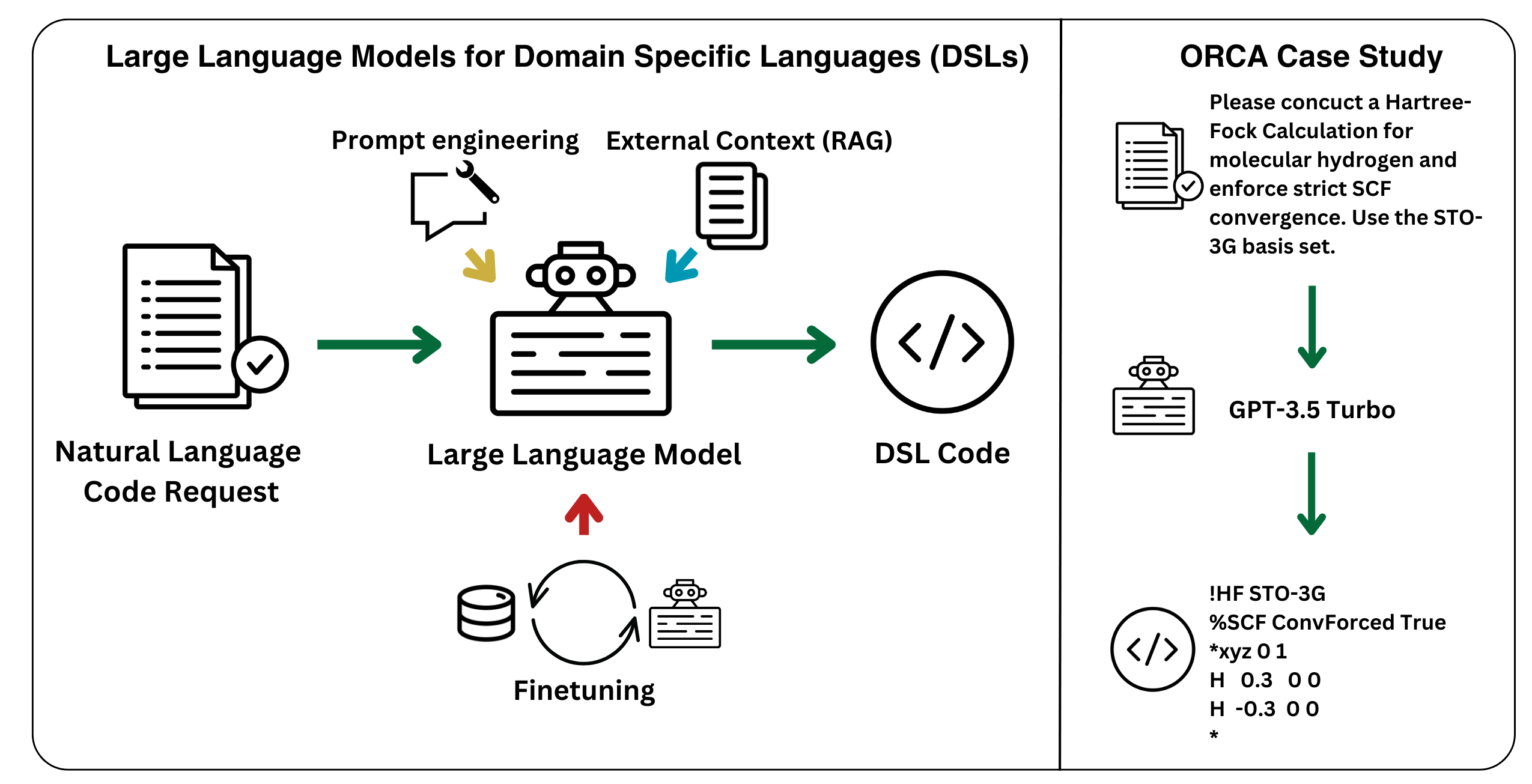
23. "Developing Large Language Models for Quantum Chemistry Simulation Input Generation"
Artificial Design
Reaction Simulation
P. F. Jacobs, R. Pollice*
ChemRxiv 2024.
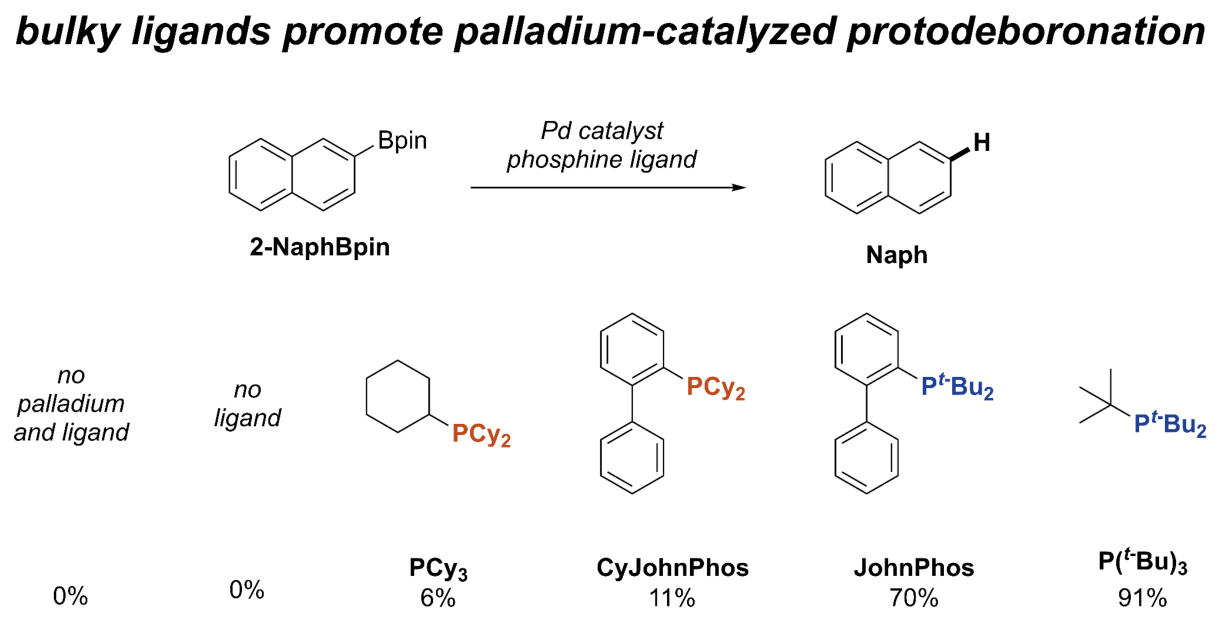
22. "Bulky phosphine ligands promote palladium-catalysed protodeboronation"
Molecular Catalysis
Reaction Simulation
C. T. Ser#, H. Hao#, S. Pablo-García, K. Jorner, S. Li, R. Pollice*, A. Aspuru-Guzik*
ChemRxiv 2024.
21. "Show, Don't Tell: Evaluating Large Language Models Beyond Textual Understanding with ChildPlay"
Artificial Design
G. Hora de Carvalho, O. Knap, R. Pollice*
arXiv 2024.

2023
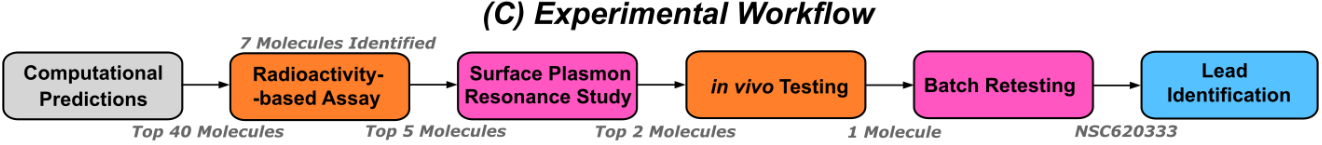
20. "Drug Discovery in Low Data Regimes: Leveraging a Computational Pipeline for the Discovery of Novel SARS-CoV-2 Nsp14-MTase Inhibitors"
Reaction Simulation
Artificial Design
A. Nigam, M. F. D. Hurley, F. Li, E. Konkoĺová, M. Klíma, J. Trylčová, R. Pollice, S. S. Çınaroğlu, R. Levin-Konigsberg, J. Handjaya, M. Schapira, I. Chau, S. Perveen, H.-L. Ng, H. Ü. Kaniskan, Y. Han, S. Singh, C. Gorgulla, A. Kundaje, J. Jin, V. A. Voelz, J. Weber, R. Nencka, E. Boura, M. Vedadi*, A. Aspuru-Guzik*
BioRxiv 2023.
19. "Ultrafast computational screening of molecules with inverted singlet-triplet energy gaps using the Pariser-Parr-Pople semi-empirical quantum chemistry method"
Reaction Simulation
K. Jorner*, R. Pollice*, C. Lavigne, A. Aspuru-Guzik*
ChemRxiv 2023.
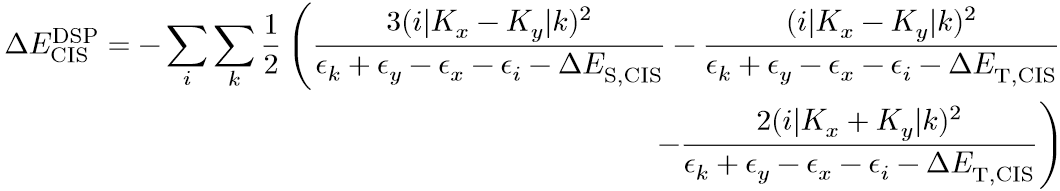
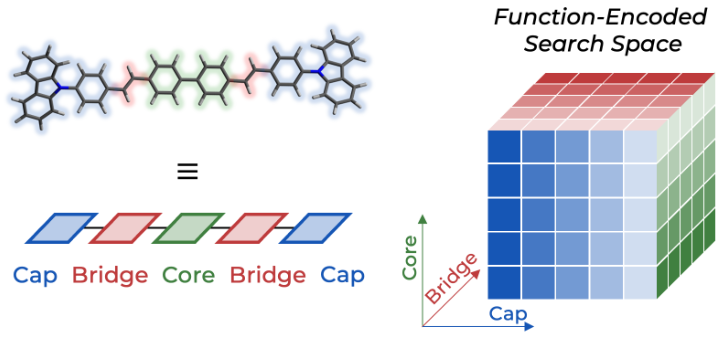
18. "Delocalized, Asynchronous, Closed-Loop Discovery of Organic Laser Emitters"
Reaction Simulation
Lab Automation
Artificial Design
F. Strieth-Kalthoff#, H. Hao#, V. Rathore, J. Derasp, T. Gaudin, N. H. Angello, M. Seifrid, E. Trushina, M. Guy, J. Liu, X. Tang, M. Mamada, W. Wang, T. Tsagaantsooj, C. Lavigne, R. Pollice, T. C. Wu, K. Hotta, L. Bodo, S. Li, M. Haddadnia, A. Wolos, R. Roszak, C. T. Ser, C. Bozal-Ginesta, R. J. Hickman, J. Vestfrid, A. Aguilar-Gránda, E. L. Klimareva, R. C. Sigerson, W. Hou, D. Gahler, S. Lach, A. Warzybok, O. Borodin, S. Rohrbach, B. Sanchez-Lengeling, C. Adachi*, B. A. Grzybowski*, L. Cronin*, J. E. Hein*, M. D. Burke*, A. Aspuru-Guzik*
ChemRxiv 2023.
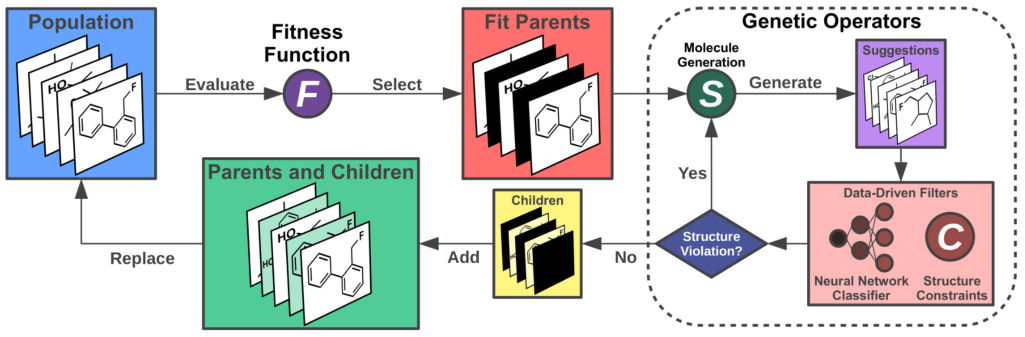
17. "Artificial Design of Organic Emitters via a Genetic Algorithm Enhanced by a Deep Neural Network"
Reaction Simulation
Artificial Design
A. Nigam#, R. Pollice#,*, P. Friederich, A. Aspuru-Guzik*
ChemRxiv 2023.
16. "Rational Design of Organic Molecules with Inverted Gaps between First Excited Singlet and Triplet"
Reaction Simulation
R. Pollice*, B. Ding, A. Aspuru-Guzik*
ChemRxiv 2023.
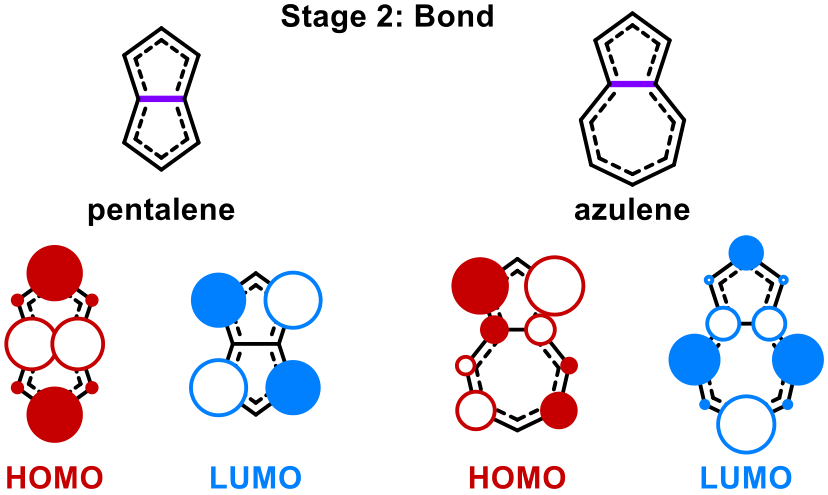
15. "On data and dimension in chemistry I - irreversibility, concealment and emergent conservation laws"
Molecular Catalysis
A. Blokhuis*, M. van Kuppeveld, D. van de Weem, R. Pollice*
arXiv 2023.
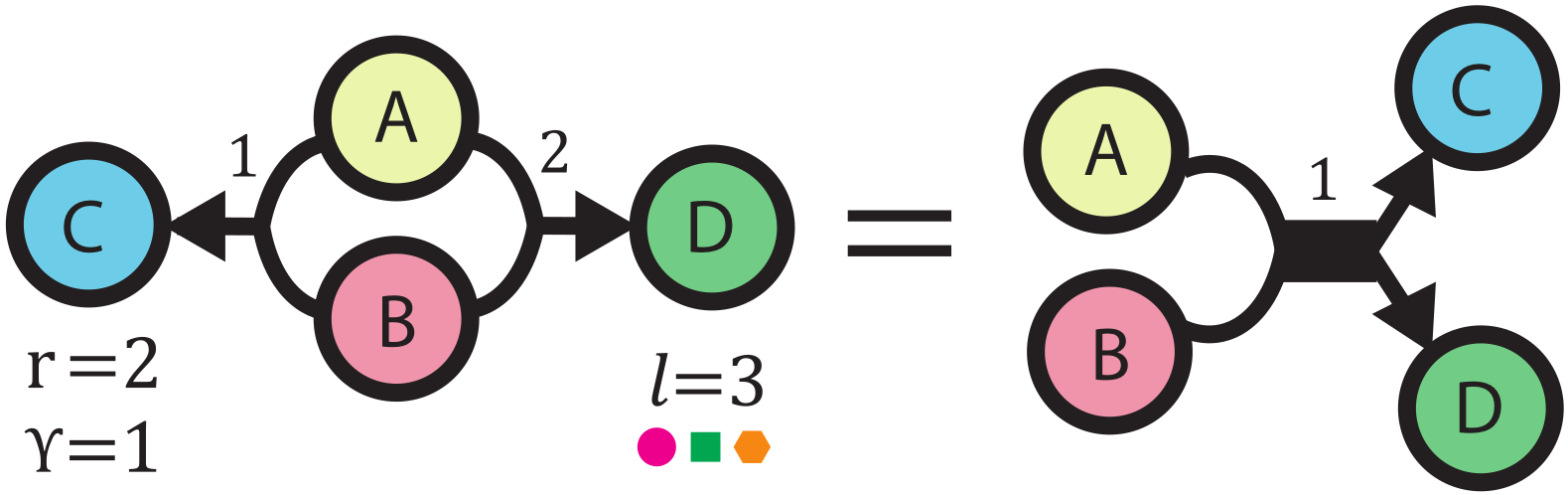
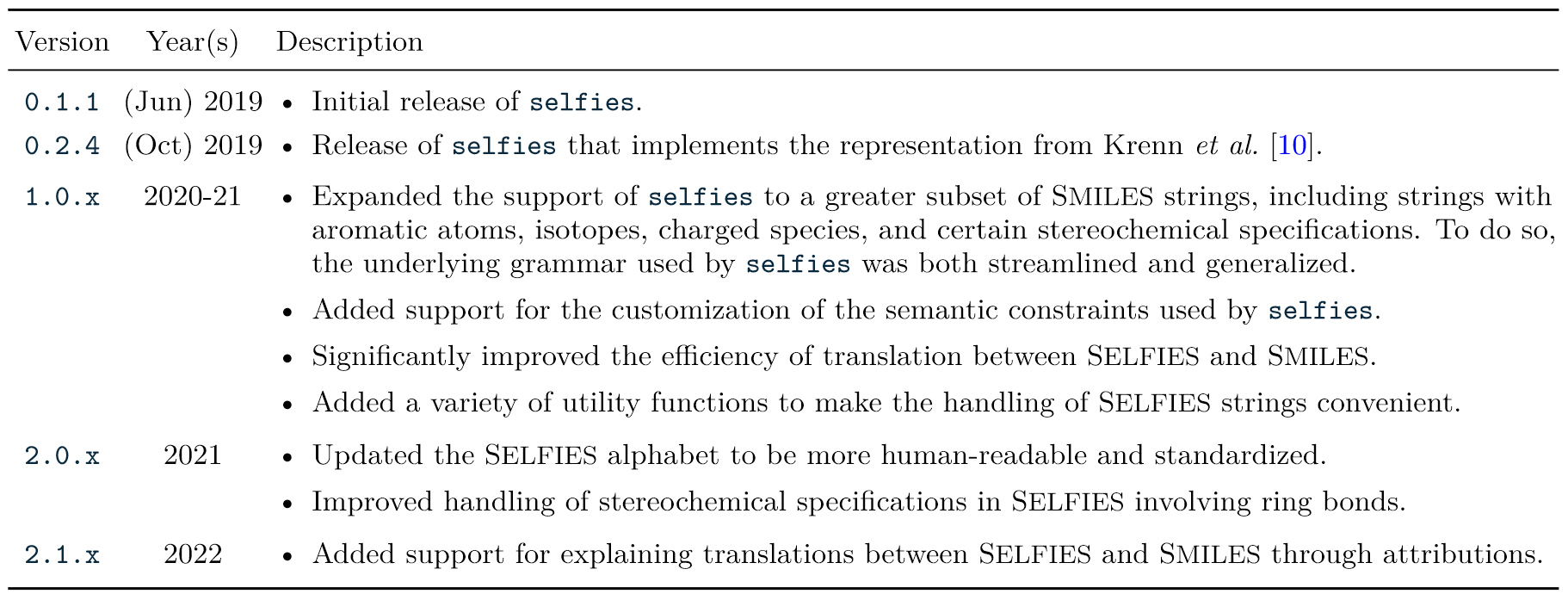
14. "Recent advances in the Self-Referencing Embedding Strings (SELFIES) library"
Artificial Design
A. Lo*, R. Pollice*, A. Nigam, A. D. White, M. Krenn, A. Aspuru-Guzik*
arXiv 2023.

2022
13. "Inverse molecular design and parameter optimization with Hückel theory using automatic differentiation"
Artificial Design
R. A. Vargas-Hernández*, K. Jorner, R. Pollice, A. Aspuru-Guzik
arXiv 2022.
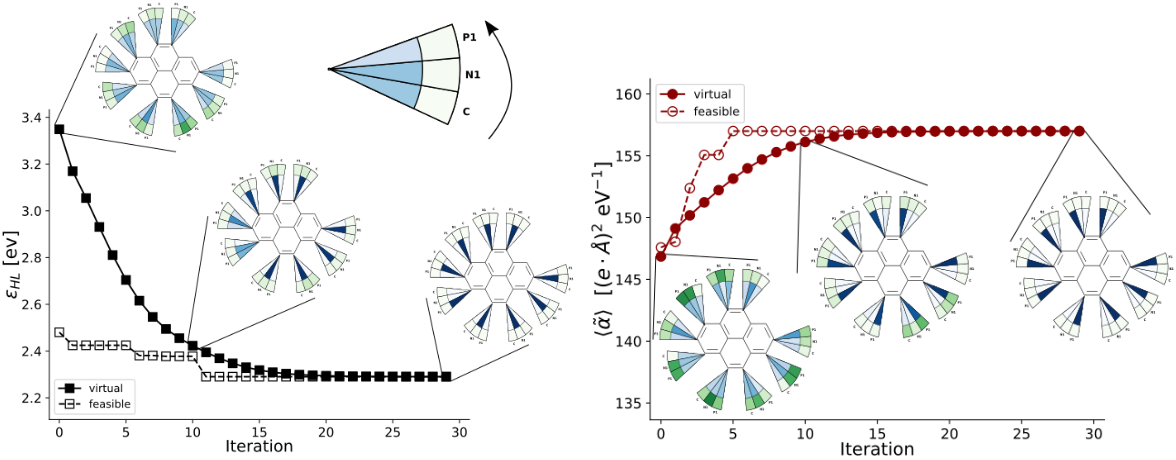
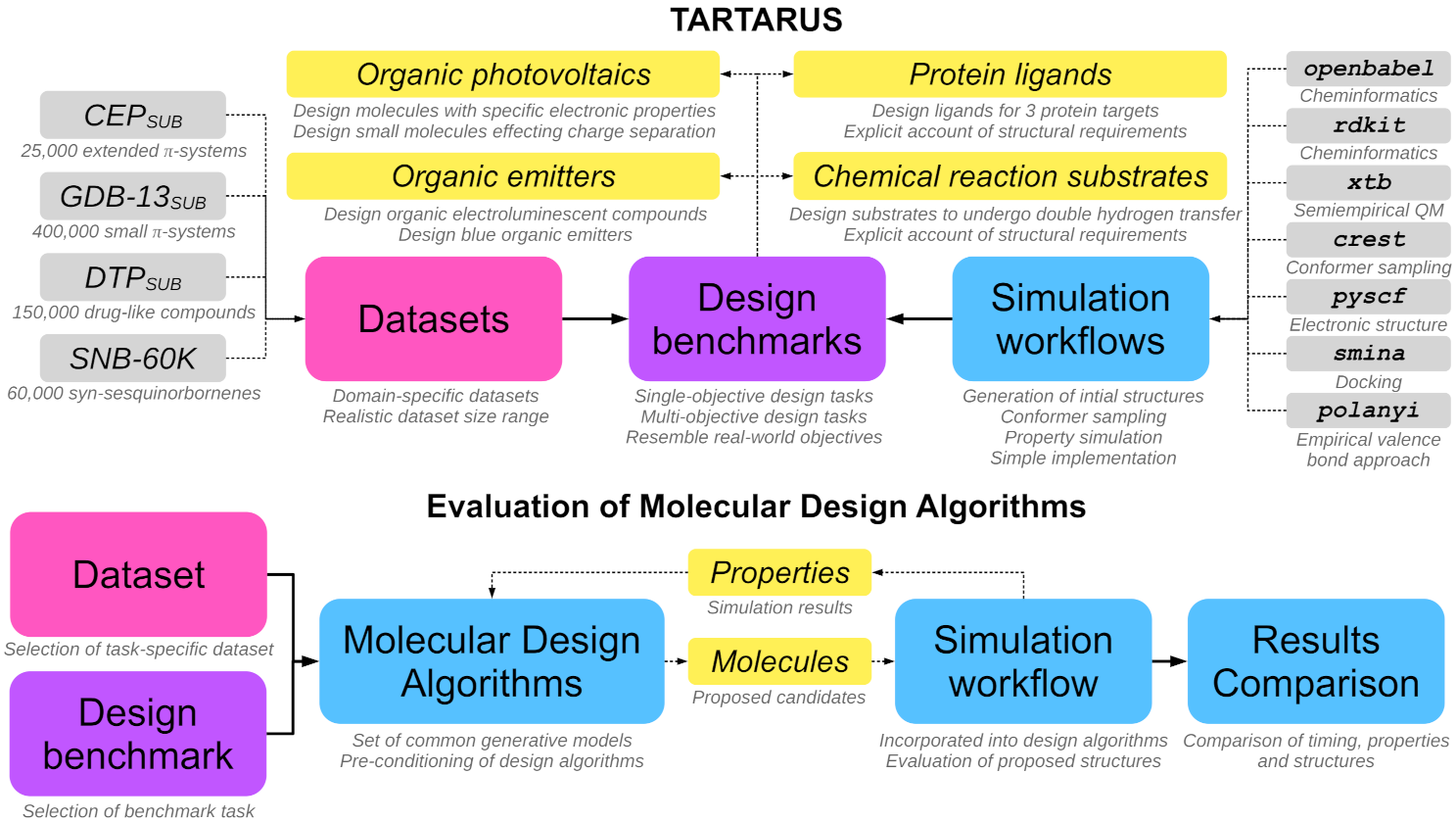
12. "Tartarus: A Benchmarking Platform for Realistic And Practical Inverse Molecular Design"
Artificial Design
A. Nigam, R. Pollice*, G. Tom, K. Jorner, L. A. Thiede, A. Kundaje, A. Aspuru-Guzik
arXiv 2022.
Prior Preprints
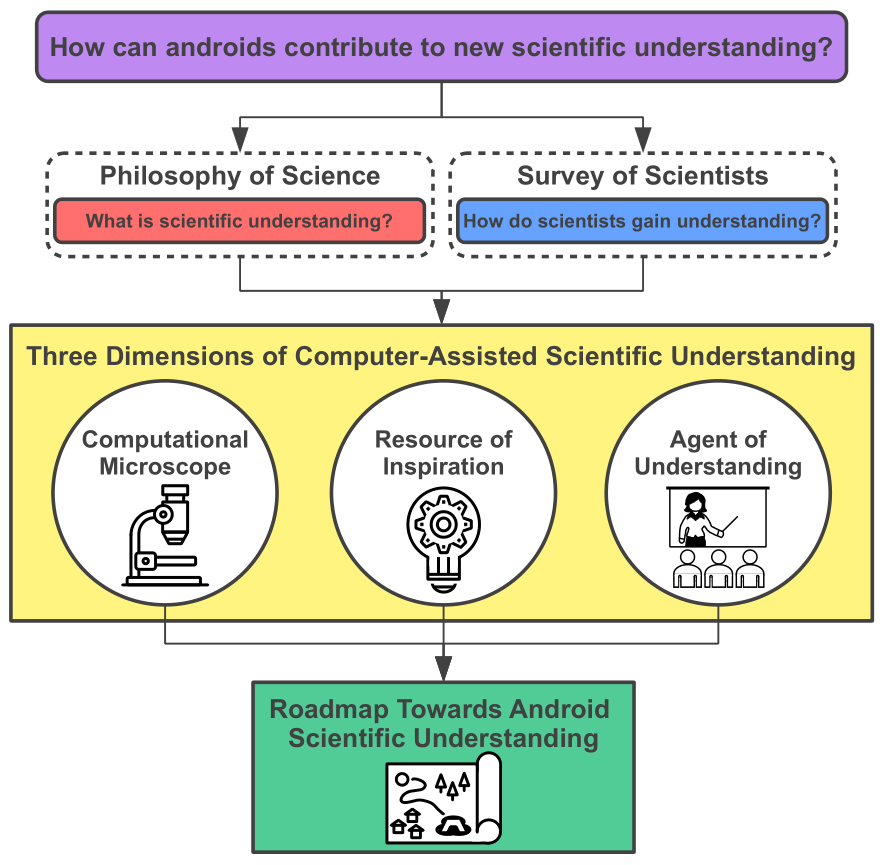
11. "On scientific understanding with artificial intelligence"
Artificial Design
M. Krenn*, R. Pollice, S. Y. Guo, M. Aldeghi, A. Cervera-Lierta, P. Friederich, G. d. P. Gomes, F. Häse, A. Jinich, A. Nigam, Z. Yao, A. Aspuru-Guzik*
arXiv 2022.
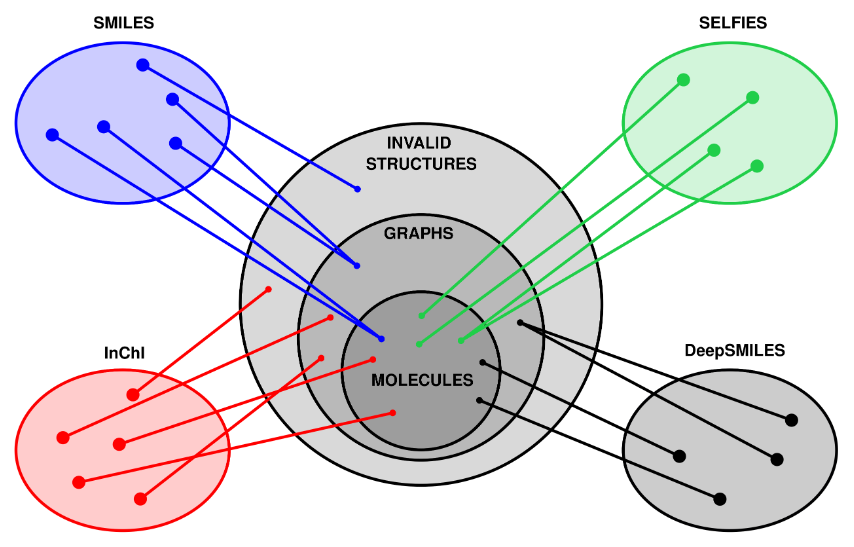
10. "SELFIES and the future of molecular string representations"
Artificial Design
Reaction Simulation
M. Krenn*, Q. Ai, S. Barthel, N. Carson, A. Frei, N. C. Frey, P. Friederich, T. Gaudin, A. A. Gayle, K. M. Jablonka, R. F. Lameiro, D. Lemm, A. Lo, S. M. Moosavi, J. M. Nápoles-Duarte, A. Nigam, R. Pollice, K. Rajan, U. Schatzschneider, P. Schwaller, M. Skreta, B. Smit, F. Strieth-Kalthoff, C. Sun, G. Tom, G. F. von Rudorff, A. Wang, A. White, A. Young, R. Yu, A. Aspuru-Guzik*
arXiv 2022.
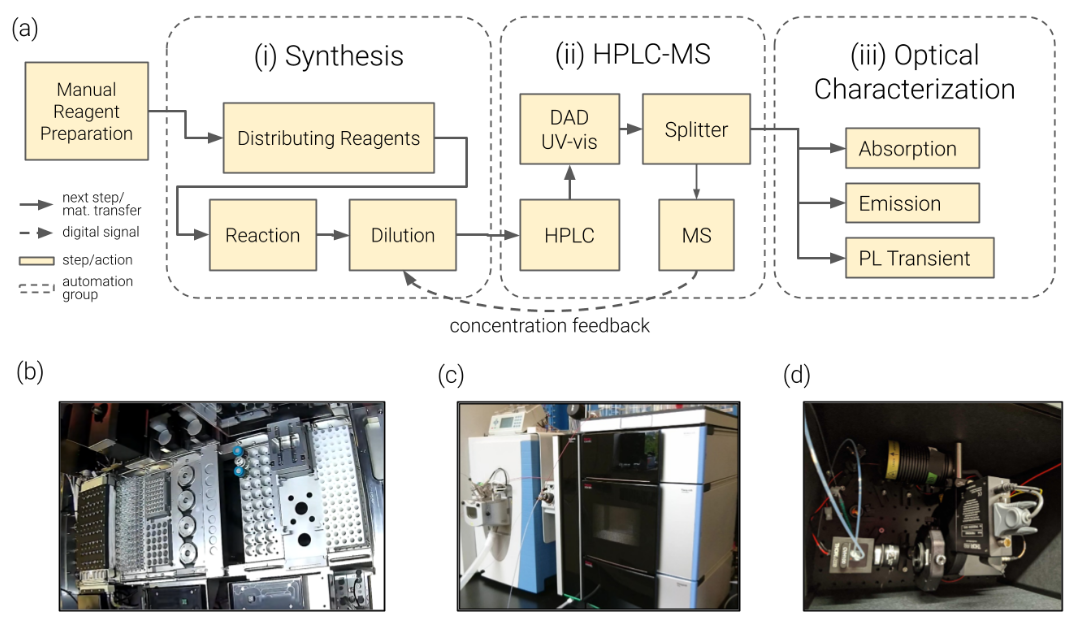
9. "A Materials Acceleration Platform for Organic Laser Discovery"
Reaction Simulation
Lab Automation
Artificial Design
T. C. Wu#, A. A. Granda#, K. Hotta, S. A. Yazdani, R. Pollice, J. Vestfrid, H. Hao, C. Lavigne, M. Seifrid, N. Angello, F. Bencheikh, J. E. Hein, M. Burke, C. Adachi, A. Aspuru-Guzik*
ChemRxiv 2022.
2021
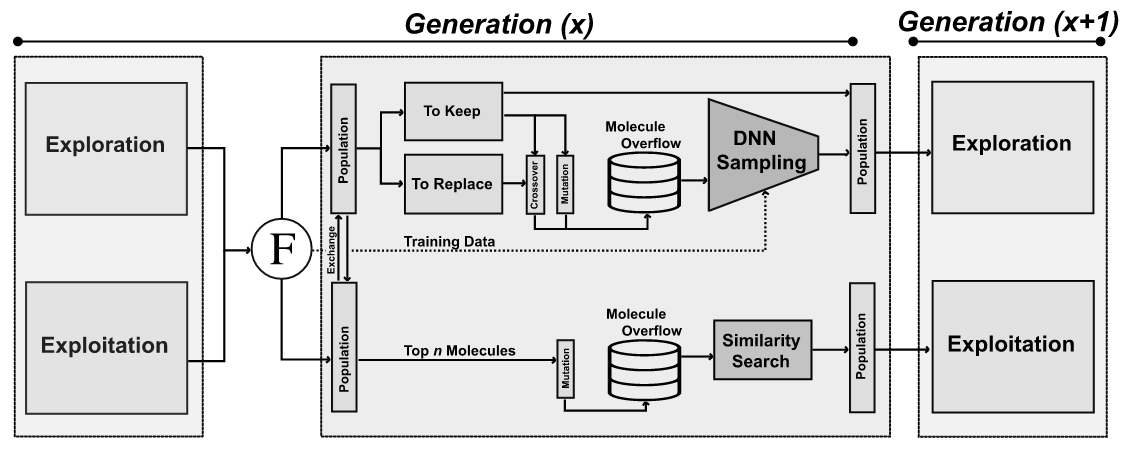
8. "JANUS: Parallel Tempered Genetic Algorithm Guided by Deep Neural Networks for Inverse Molecular Design"
Artificial Design
A. Nigam#, R. Pollice#, A. Aspuru-Guzik*
arXiv 2021.
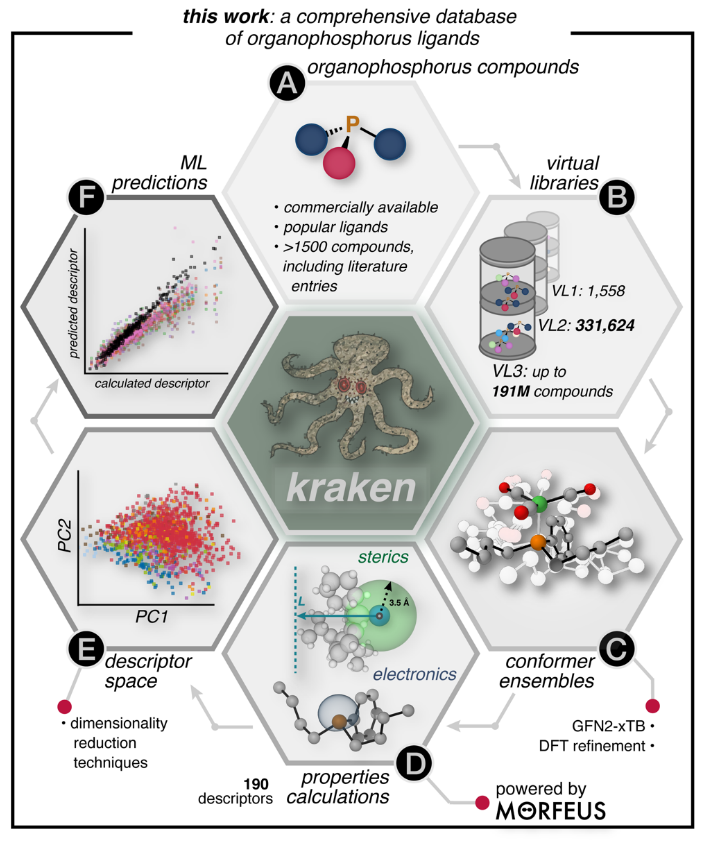
7. "A Comprehensive Discovery Platform for Organophosphorus Ligands for Catalysis"
Molecular Catalysis
Reaction Simulation
Artificial Design
T. Gensch#, G. d. P. Gomes#, P. Friederich#, E. Peters, T. Gaudin, R. Pollice, K. Jorner, A. Nigam, M. Lindner-D'Addario, M. S. Sigman*, A. Aspuru-Guzik*
ChemRxiv 2021.
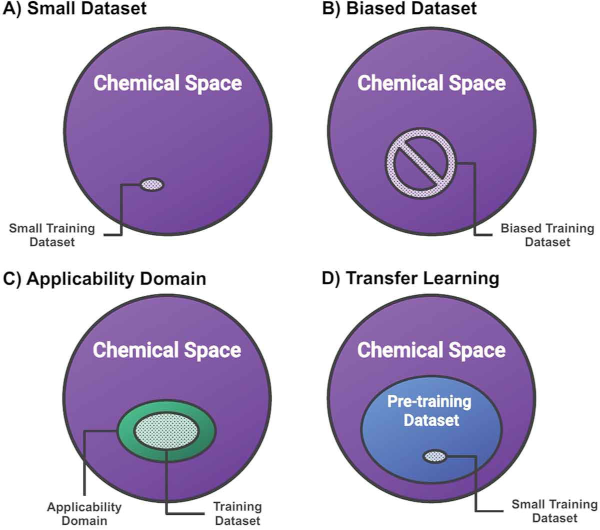
6. "Assigning Confidence to Molecular Property Prediction"
Artificial Design
Reaction Simulation
A. Nigam, R. Pollice, M. F. D. Hurley, R. J. Hickman, M. Aldeghi, N. Yoshikawa, S. Chithrananda, V. A. Voelz, A. Aspuru-Guzik*
arXiv 2021.
2020
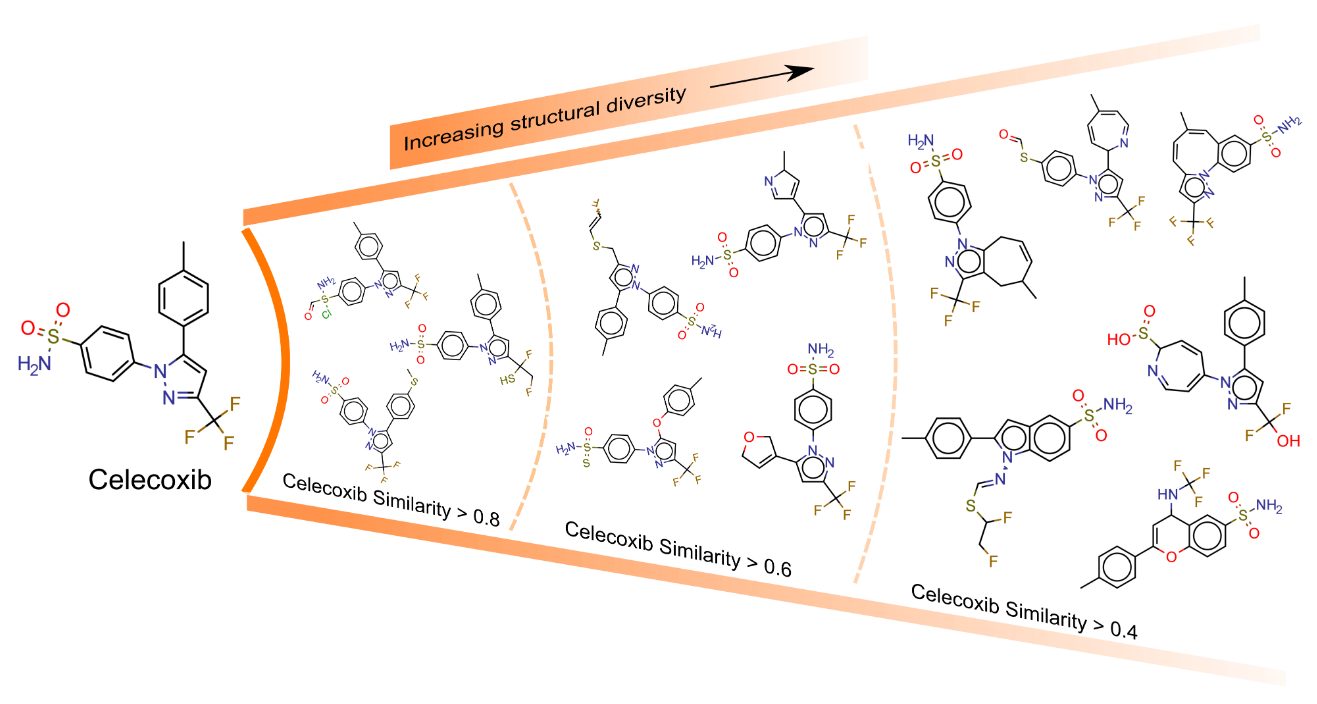
5. "Beyond Generative Models: Superfast Traversal, Optimization, Novelty, Exploration and Discovery (STONED) Algorithm for Molecules using SELFIES"
Artificial Design
A. Nigam, R. Pollice, M. Krenn, G. d. P. Gomes, A. Aspuru-Guzik*
ChemRxiv 2020.
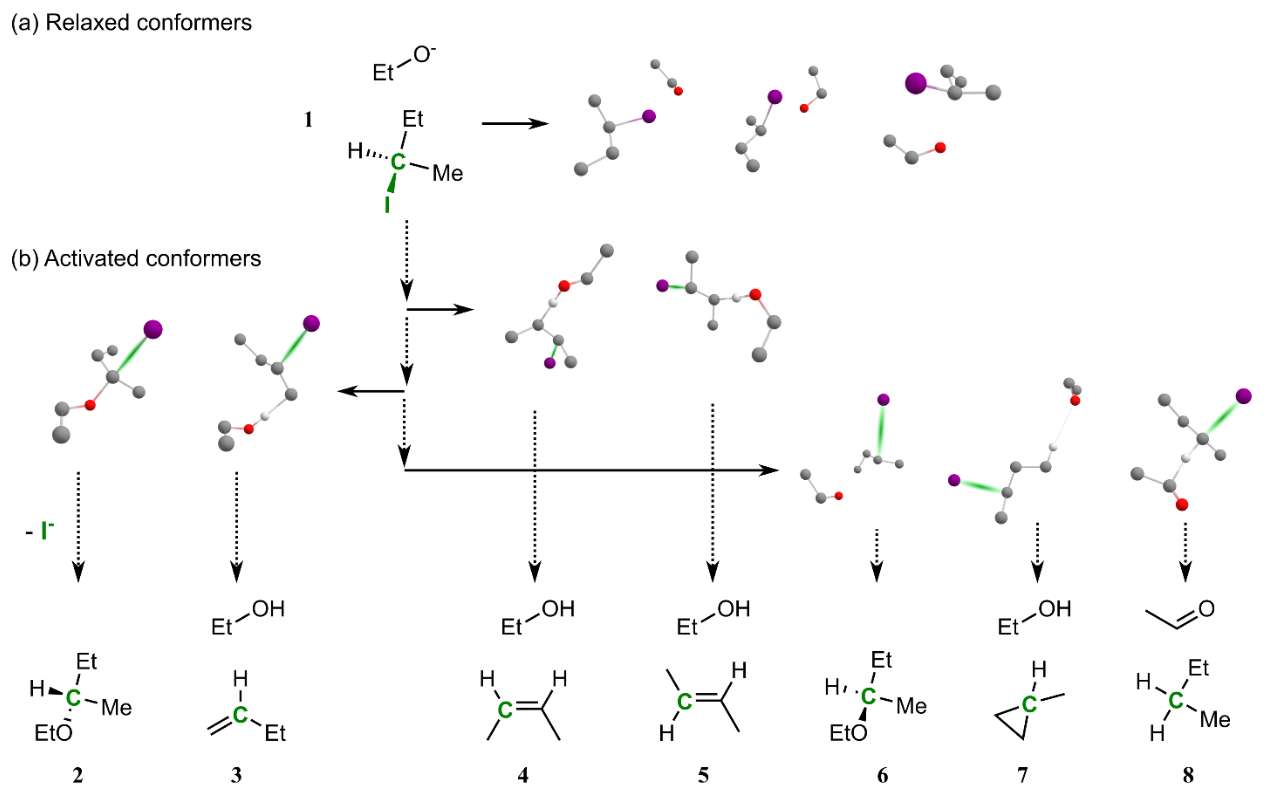
4. "Automatic discovery of chemical reactions using imposed activation"
Reaction Simulation
C. Lavigne, G. d. P. Gomes#,*, R. Pollice#,*, A. Aspuru-Guzik*
ChemRxiv 2020.
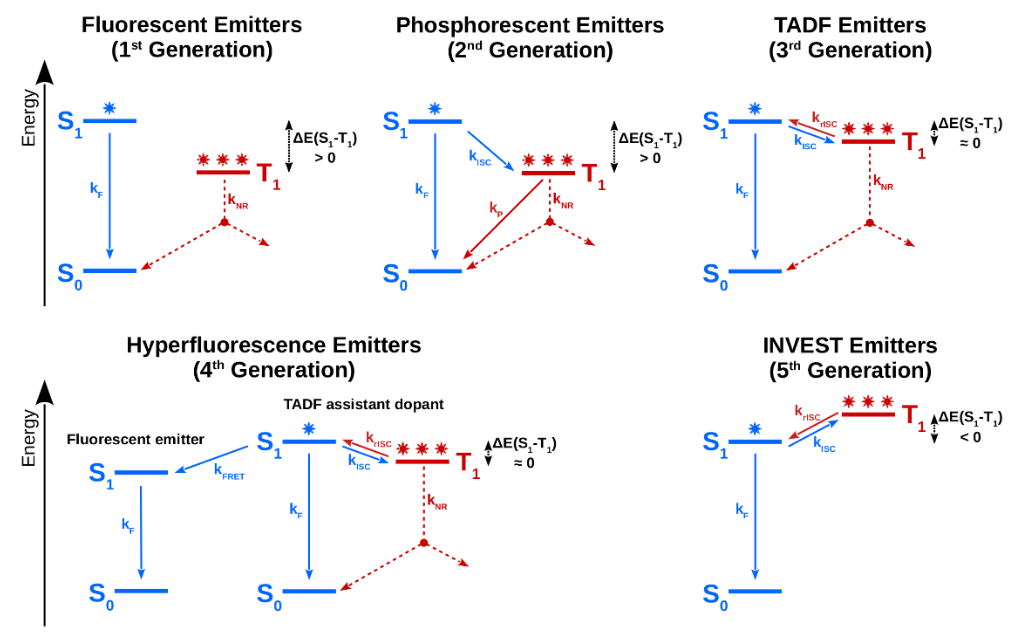
3. "Organic Molecules with Inverted Gaps between First Excited Singlet and Triplet States and Appreciable Fluorescence Rates"
Reaction Simulation
R. Pollice, P. Friederich, C. Lavigne, G. d. P. Gomes, A. Aspuru-Guzik*
ChemRxiv 2020.
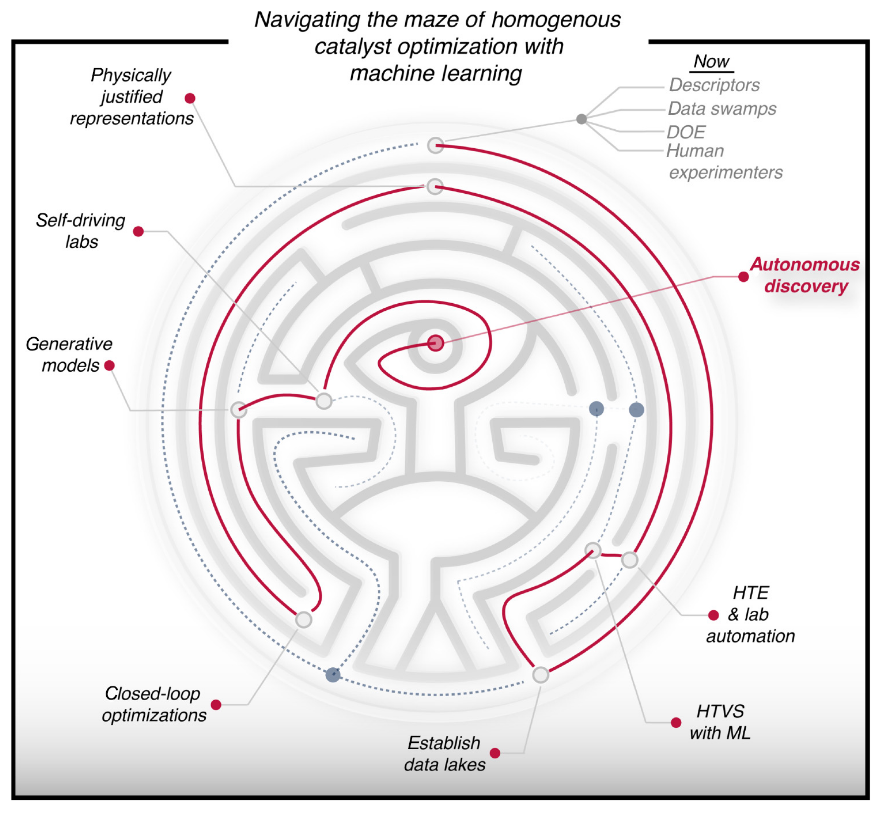
2. "Navigating through the Maze of Homogeneous Catalyst Design with Machine Learning"
Molecular Catalysis
Reaction Simulation
Lab Automation
Artificial Design
G. d. P. Gomes#, R. Pollice#, A. Aspuru-Guzik*
ChemRxiv 2020.
2019
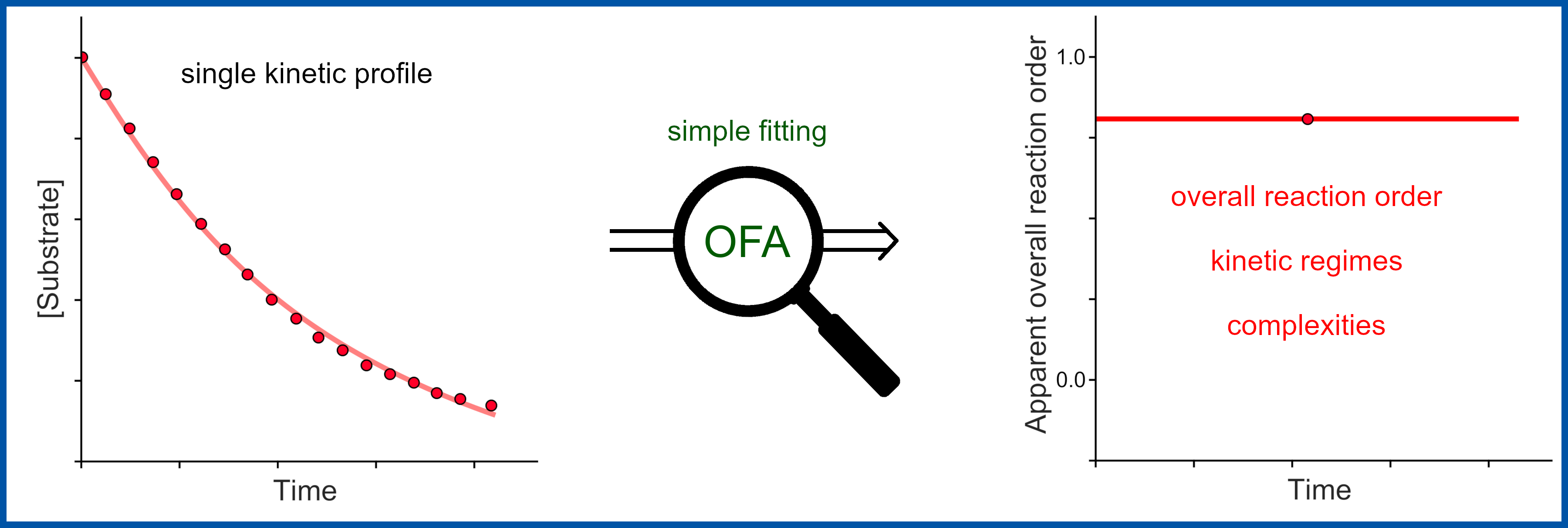
1. "A General Fitting Function to Estimate Apparent Reaction Orders of Kinetic Profiles"
Molecular Catalysis
R. Pollice*
ChemRxiv 2019.